Summary
Globals
| Reference size | 2,240,221,656 |
| Number of reads | 28,590,284 |
| Mapped reads | 28,590,284 / 100% |
| Unmapped reads | 0 / 0% |
| Mapped paired reads | 28,590,284 / 100% |
| Mapped reads, first in pair | 14,295,142 / 50% |
| Mapped reads, second in pair | 14,295,142 / 50% |
| Mapped reads, both in pair | 28,590,284 / 100% |
| Mapped reads, singletons | 0 / 0% |
| Read min/max/mean length | 20 / 145 / 99.78 |
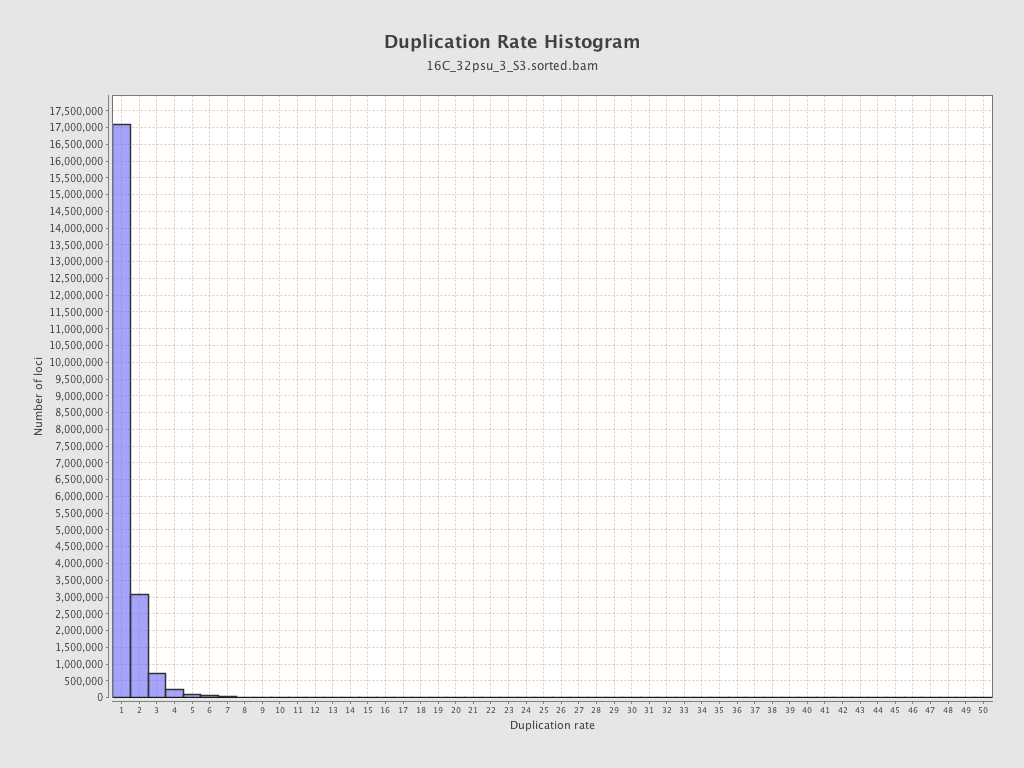
| Duplicated reads (estimated) | 7,200,829 / 25.19% |
| Duplication rate | 20.03% |
| Clipped reads | 0 / 0% |
ACGT Content
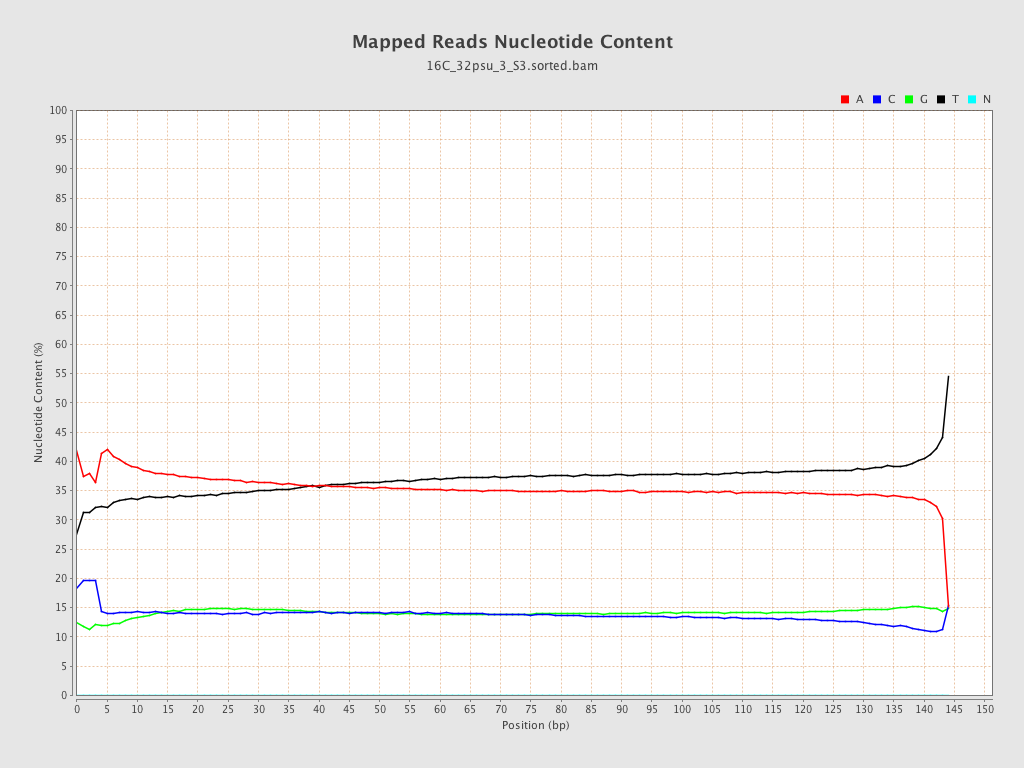
| Number/percentage of A's | 1,020,527,321 / 36.02% |
| Number/percentage of C's | 396,941,658 / 14.01% |
| Number/percentage of T's | 1,019,767,148 / 35.99% |
| Number/percentage of G's | 396,242,508 / 13.98% |
| Number/percentage of N's | 17,255 / 0% |
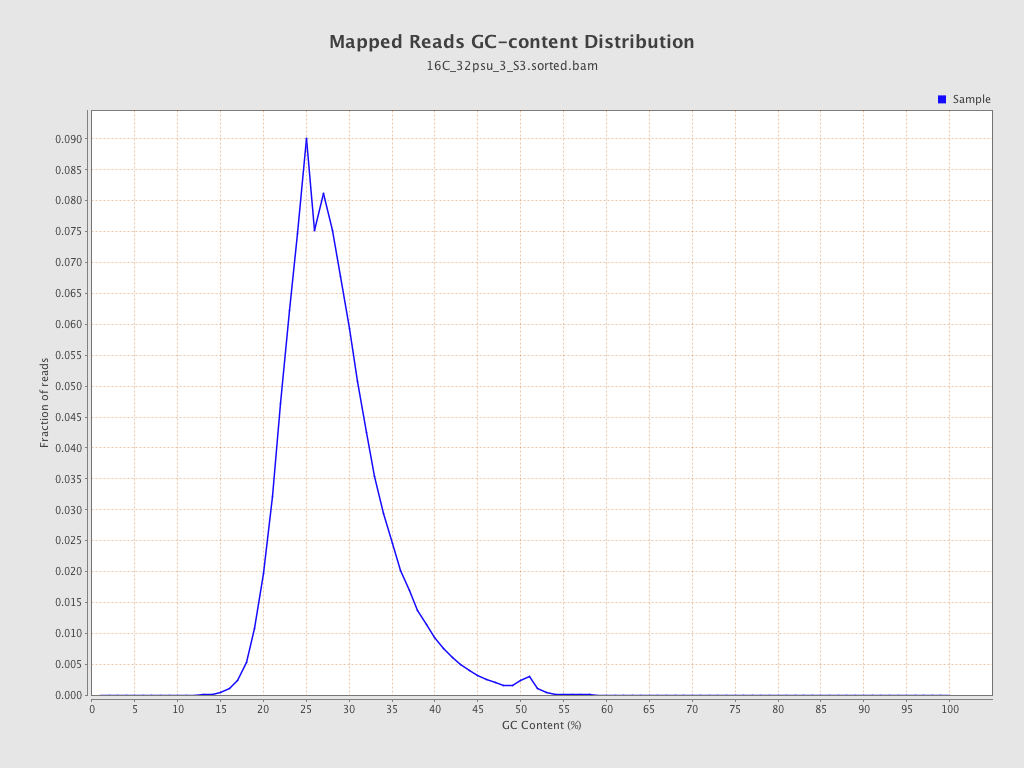
| GC Percentage | 27.99% |
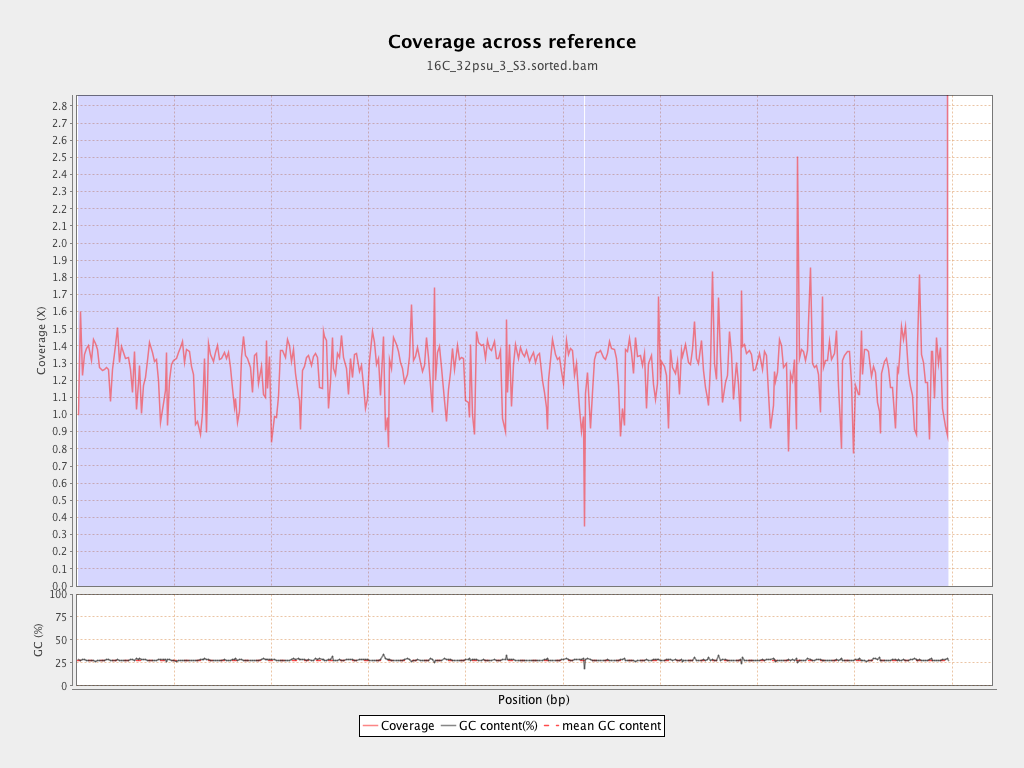
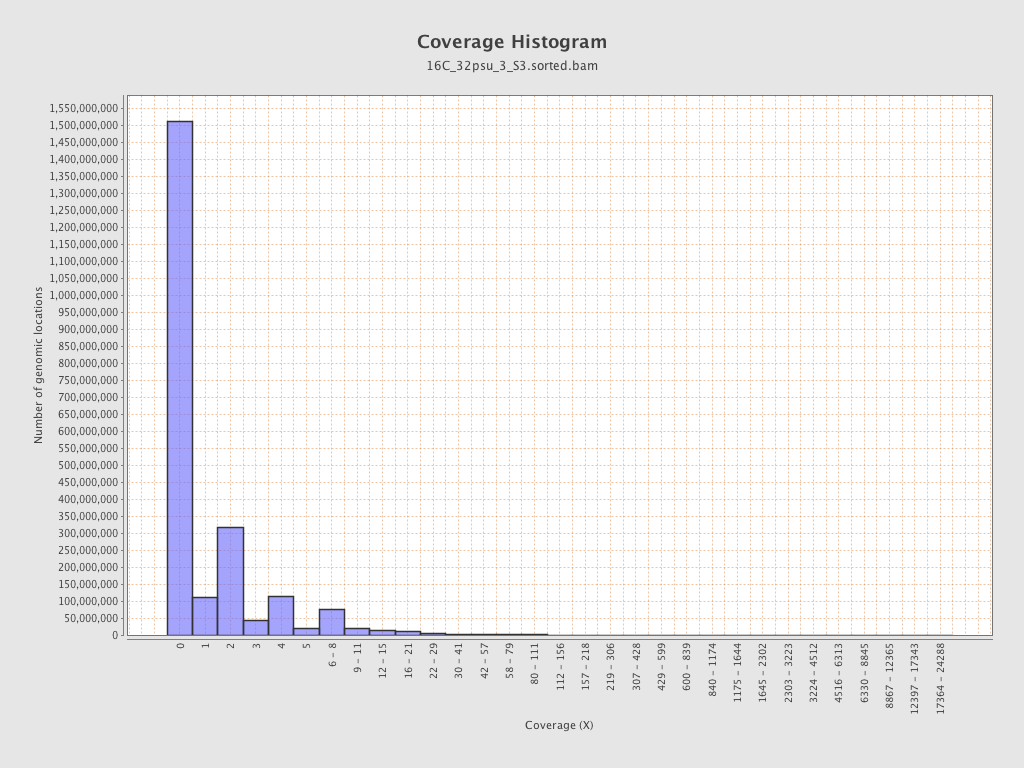
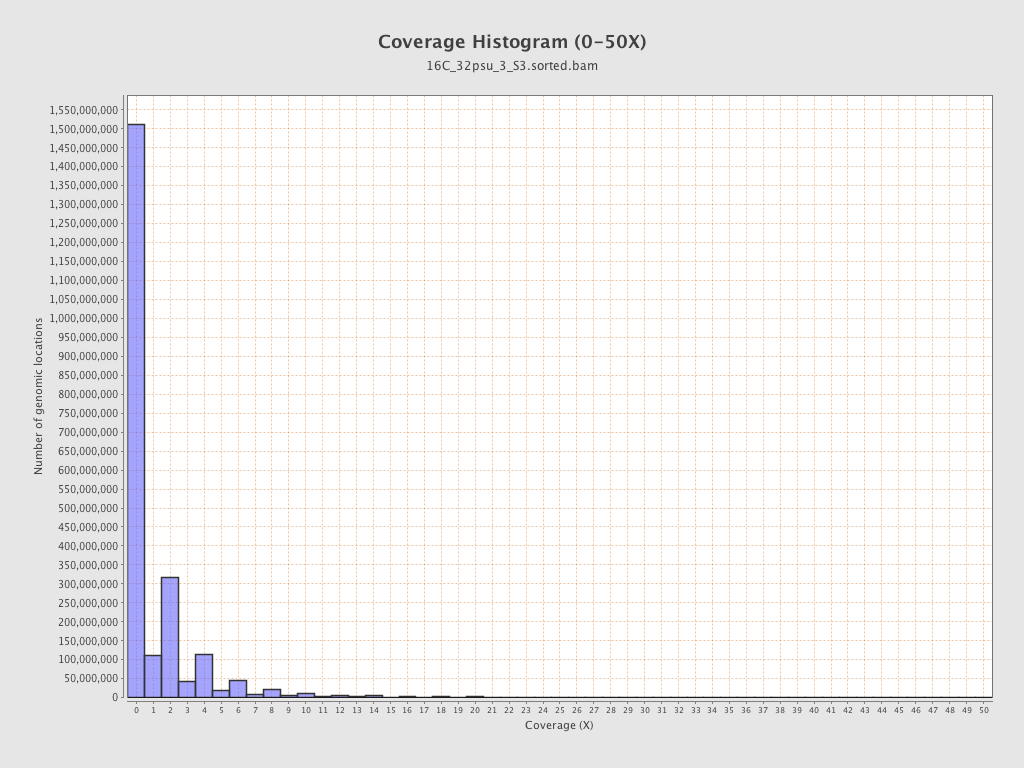
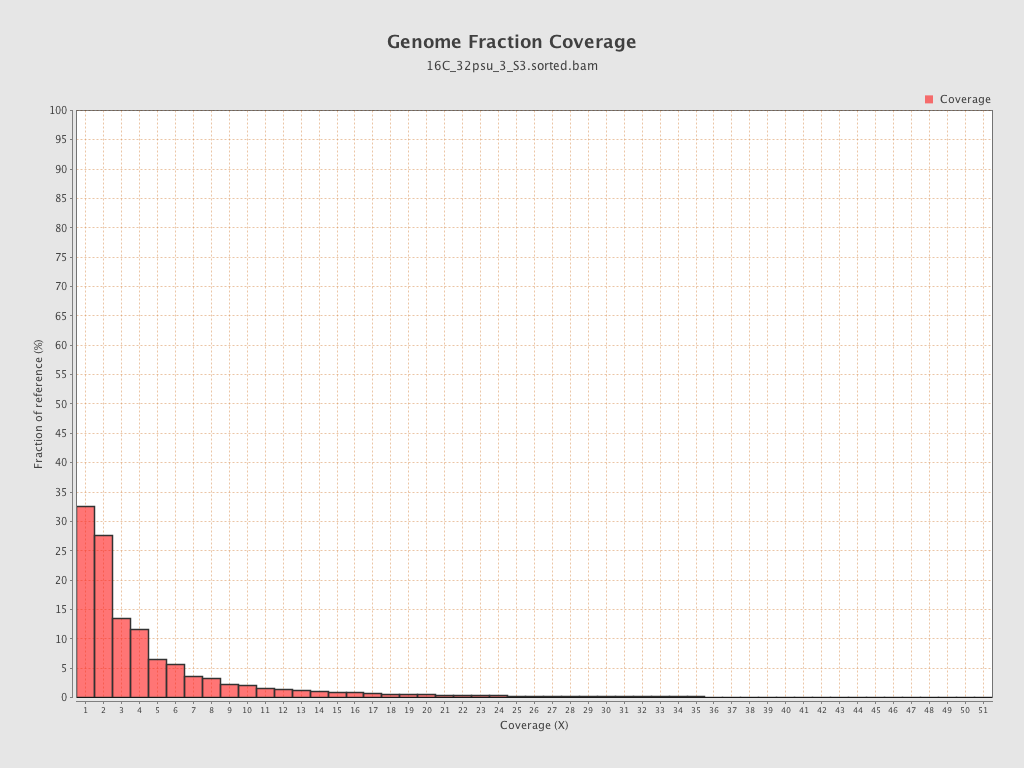
Coverage
| Mean | 1.2682 |
| Standard Deviation | 10.7154 |
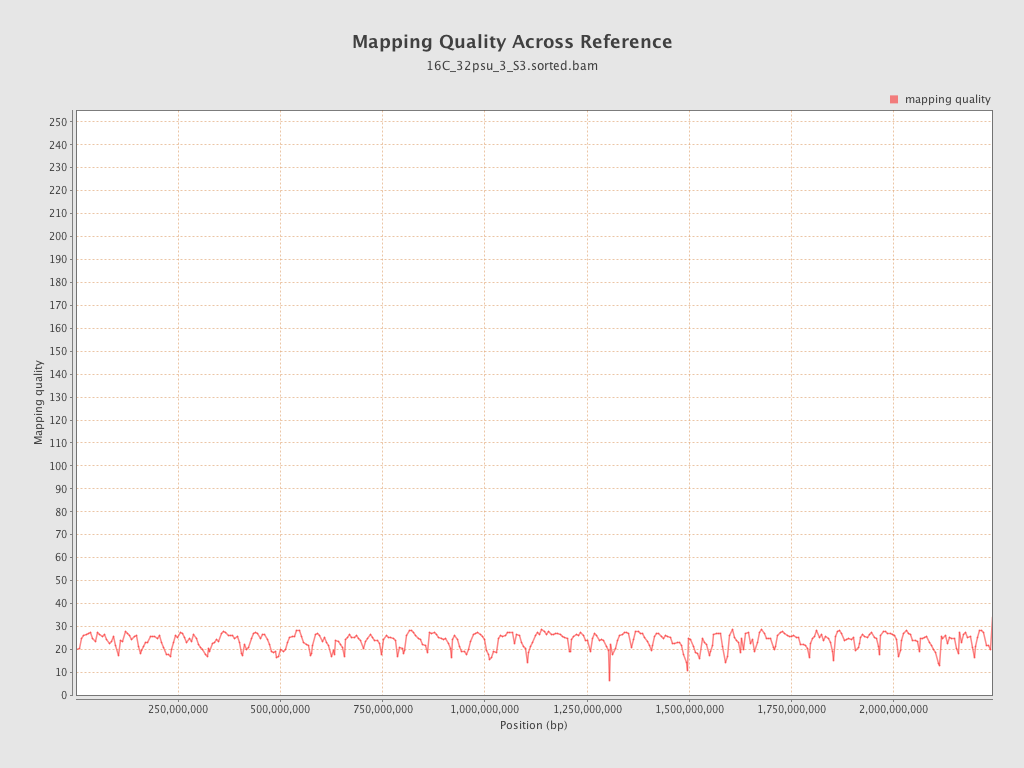
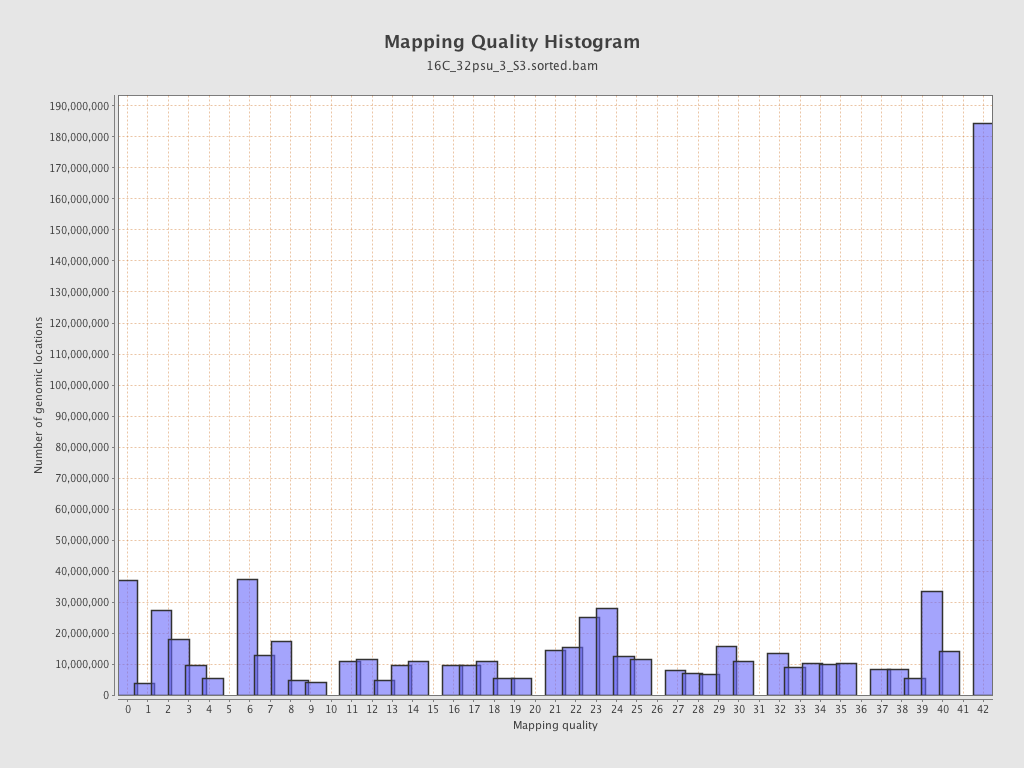
Mapping Quality
| Mean Mapping Quality | 23.52 |
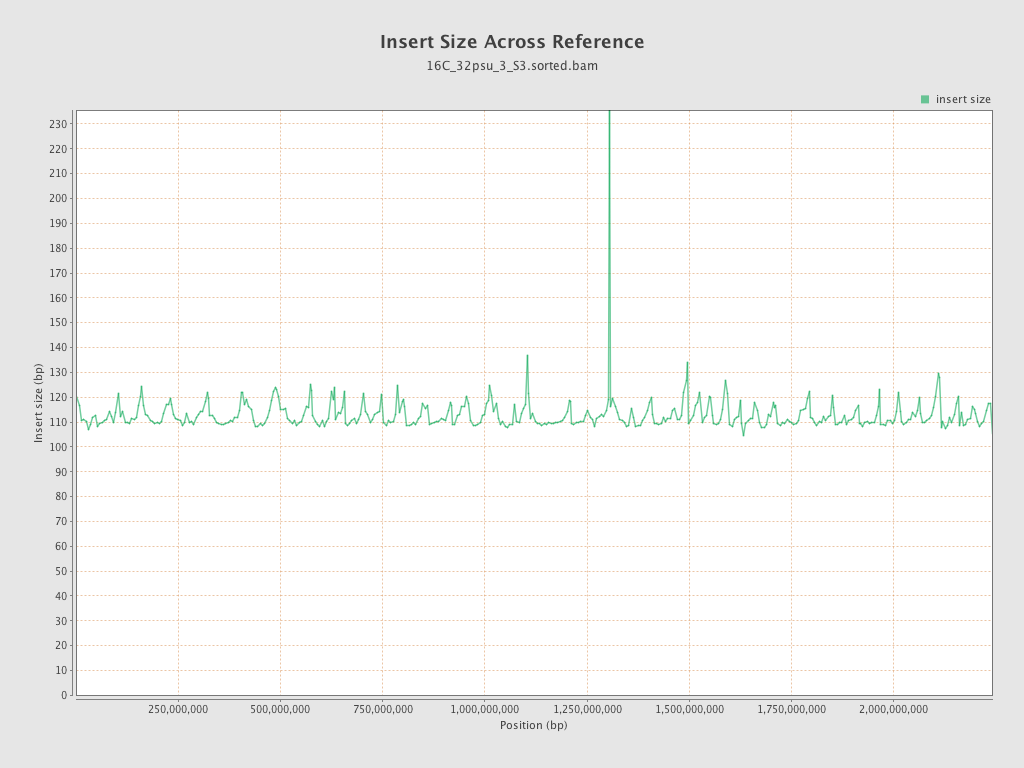
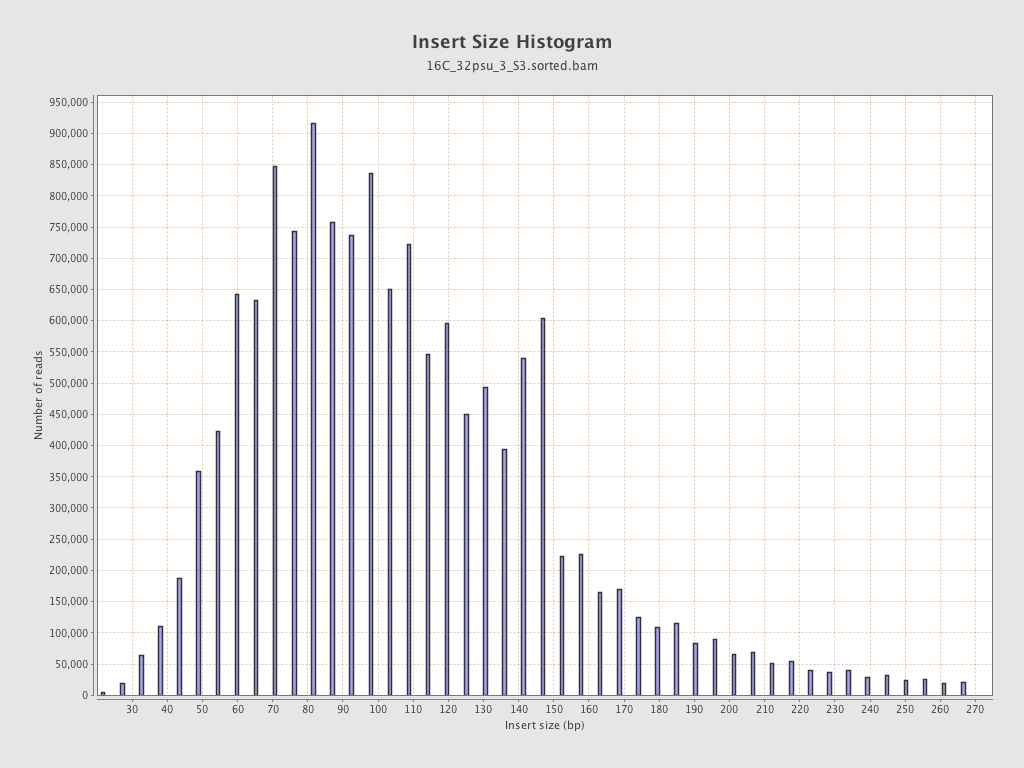
Insert size
| Mean | 112.61 |
| Standard Deviation | 51.19 |
| P25/Median/P75 | 78 / 103 / 136 |
Mismatches and indels
| General error rate | 20.97% |
| Mismatches | 575,497,392 |
| Insertions | 6,578,592 |
| Mapped reads with at least one insertion | 17.02% |
| Deletions | 4,585,598 |
| Mapped reads with at least one deletion | 13.83% |
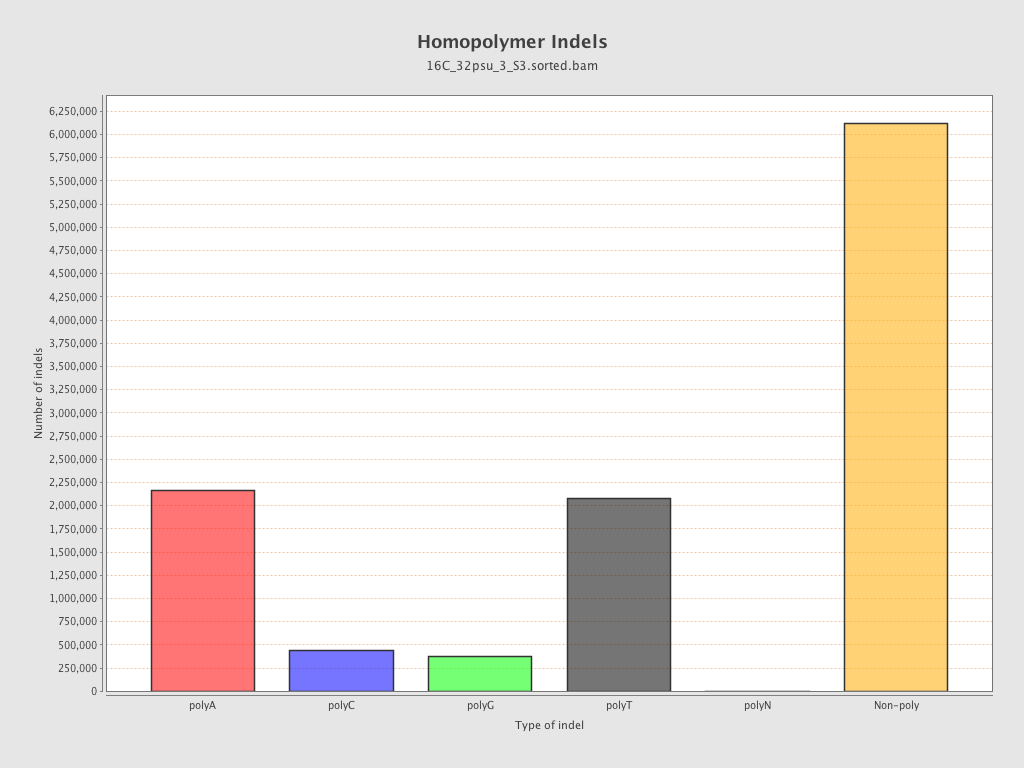
| Homopolymer indels | 45.22% |
Chromosome stats
| Name | Length | Mapped bases | Mean coverage | Standard deviation |
| NC_027300.1 | 159038749 | 207715709 | 1.3061 | 5.6386 |
| NC_027301.1 | 72943711 | 88696168 | 1.216 | 4.5431 |
| NC_027302.1 | 92503428 | 112845984 | 1.2199 | 3.972 |
| NC_027303.1 | 82398023 | 106033471 | 1.2868 | 9.4695 |
| NC_027304.1 | 80503876 | 98761548 | 1.2268 | 6.5793 |
| NC_027305.1 | 87043187 | 106183262 | 1.2199 | 8.4724 |
| NC_027306.1 | 58785265 | 74644098 | 1.2698 | 7.9122 |
| NC_027307.1 | 26434011 | 34447628 | 1.3032 | 14.4951 |
| NC_027308.1 | 141712163 | 178187998 | 1.2574 | 7.7346 |
| NC_027309.1 | 116138521 | 153122194 | 1.3184 | 15.9166 |
| NC_027310.1 | 93888508 | 116157430 | 1.2372 | 5.3317 |
| NC_027311.1 | 91880962 | 117811775 | 1.2822 | 10.6791 |
| NC_027312.1 | 107758822 | 138273214 | 1.2832 | 5.1711 |
| NC_027313.1 | 93901823 | 118284209 | 1.2597 | 7.719 |
| NC_027314.1 | 103963436 | 128635495 | 1.2373 | 4.4004 |
| NC_027315.1 | 87796322 | 113789962 | 1.2961 | 6.1235 |
| NC_027316.1 | 57682537 | 71163723 | 1.2337 | 5.802 |
| NC_027317.1 | 70695409 | 93212208 | 1.3185 | 8.4086 |
| NC_027318.1 | 82978132 | 107914289 | 1.3005 | 31.8303 |
| NC_027319.1 | 86795997 | 110795767 | 1.2765 | 7.9889 |
| NC_027320.1 | 58021487 | 70583350 | 1.2165 | 3.9431 |
| NC_027321.1 | 63420196 | 86305290 | 1.3608 | 24.1835 |
| NC_027322.1 | 49854004 | 64124643 | 1.2862 | 5.4563 |
| NC_027323.1 | 48650976 | 58254212 | 1.1974 | 5.4907 |
| NC_027324.1 | 51481326 | 64435123 | 1.2516 | 6.9037 |
| NC_027325.1 | 47900953 | 57800237 | 1.2067 | 5.6665 |
| NC_027326.1 | 43943985 | 55260800 | 1.2575 | 5.8788 |
| NC_027327.1 | 39600944 | 51445465 | 1.2991 | 12.0378 |
| NC_027328.1 | 42488238 | 49715643 | 1.1701 | 6.6333 |
| NC_001960.1 | 16665 | 6404358 | 384.2999 | 545.8597 |