Summary
Globals
| Reference size | 2,240,221,656 |
| Number of reads | 39,383,180 |
| Mapped reads | 39,383,180 / 100% |
| Unmapped reads | 0 / 0% |
| Mapped paired reads | 39,383,180 / 100% |
| Mapped reads, first in pair | 19,691,590 / 50% |
| Mapped reads, second in pair | 19,691,590 / 50% |
| Mapped reads, both in pair | 39,383,180 / 100% |
| Mapped reads, singletons | 0 / 0% |
| Read min/max/mean length | 20 / 145 / 100.88 |
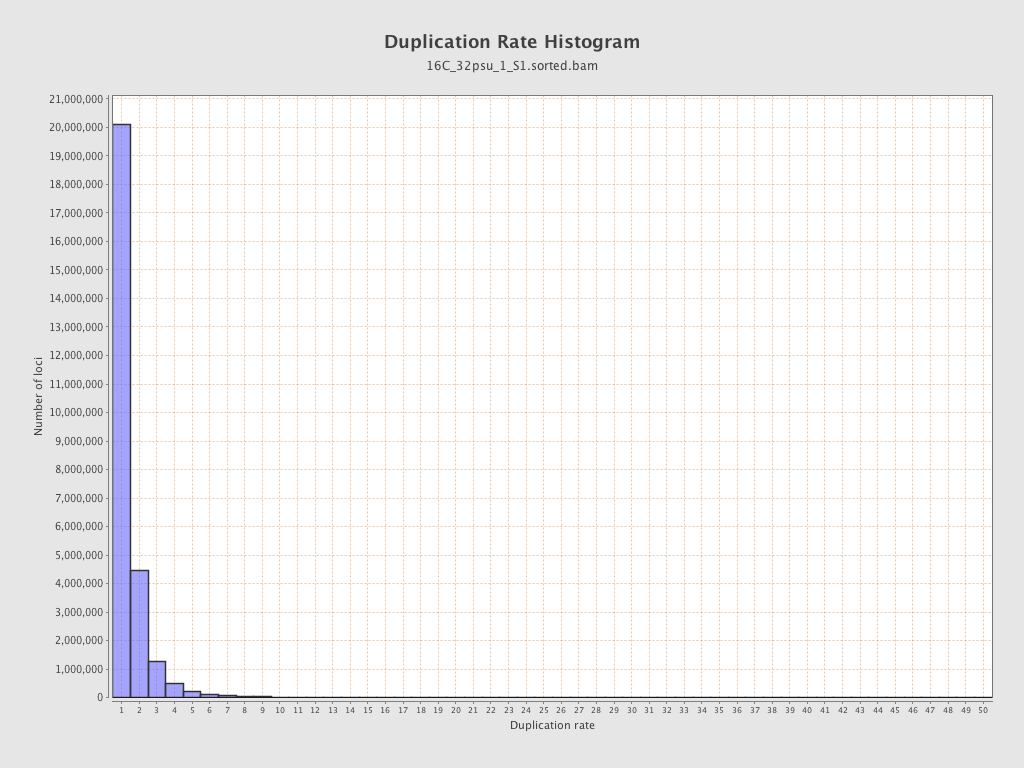
| Duplicated reads (estimated) | 12,512,260 / 31.77% |
| Duplication rate | 25.12% |
| Clipped reads | 0 / 0% |
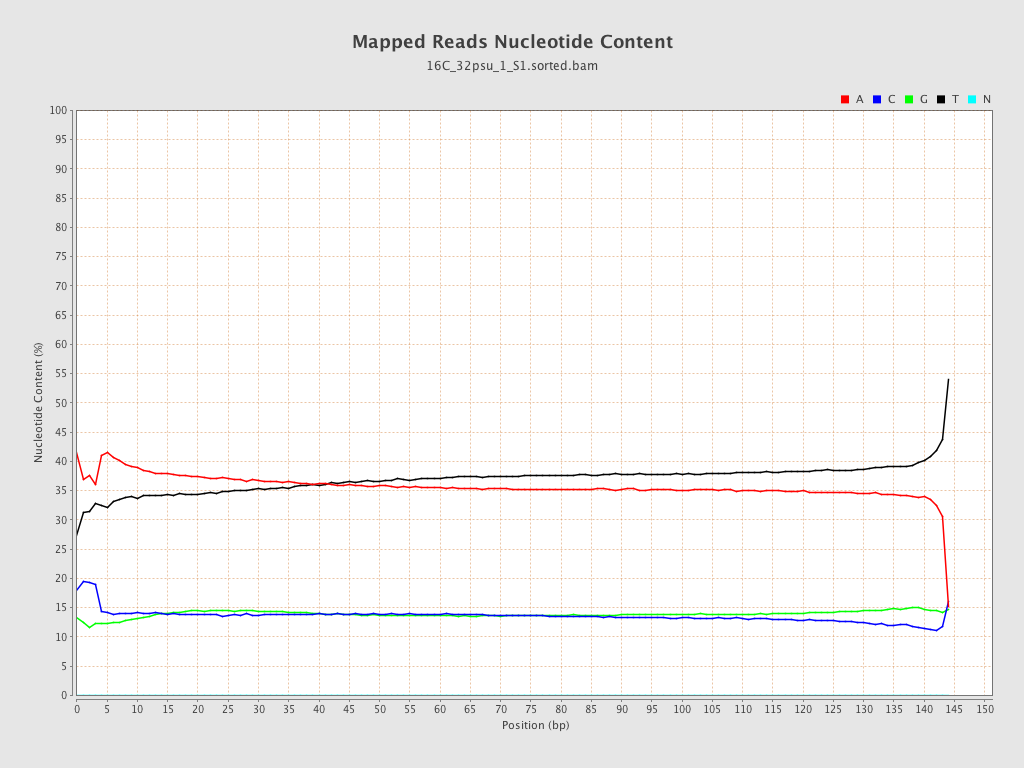
ACGT Content
| Number/percentage of A's | 1,428,843,411 / 36.22% |
| Number/percentage of C's | 544,870,442 / 13.81% |
| Number/percentage of T's | 1,428,060,144 / 36.2% |
| Number/percentage of G's | 543,297,017 / 13.77% |
| Number/percentage of N's | 24,416 / 0% |
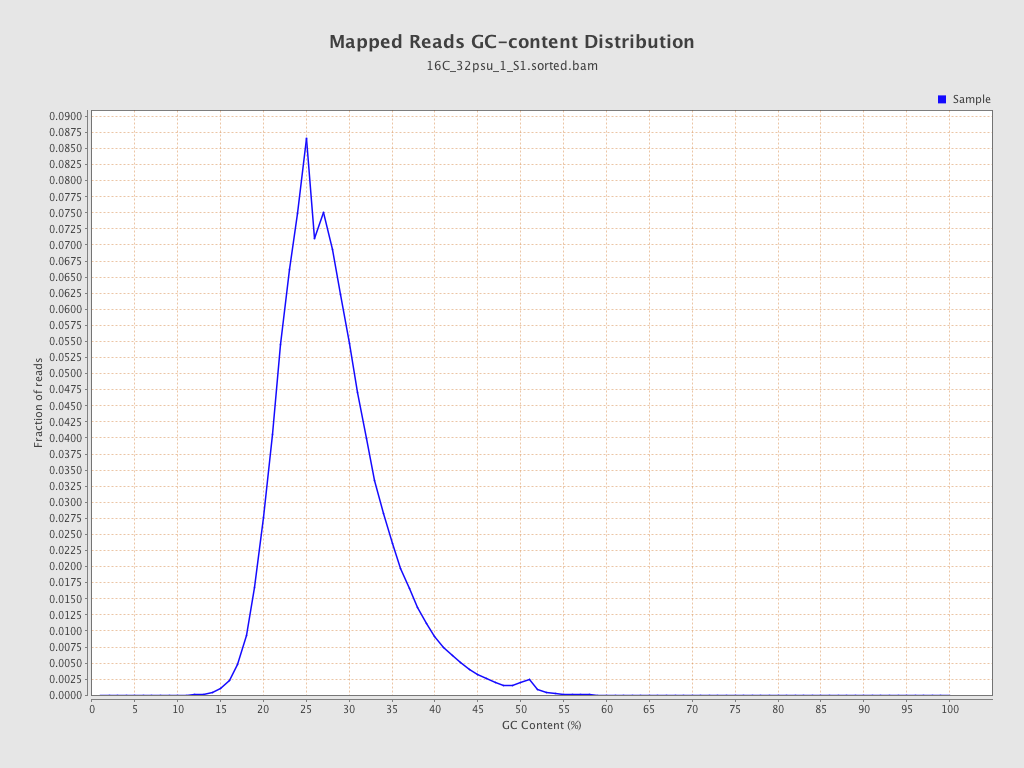
| GC Percentage | 27.58% |
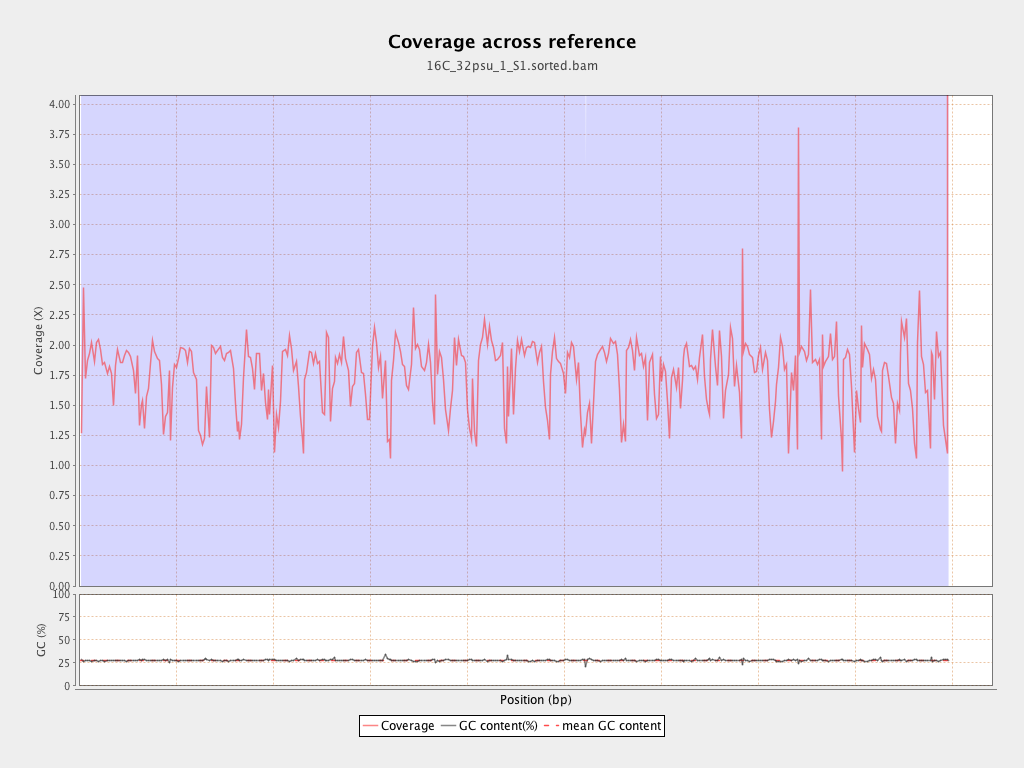
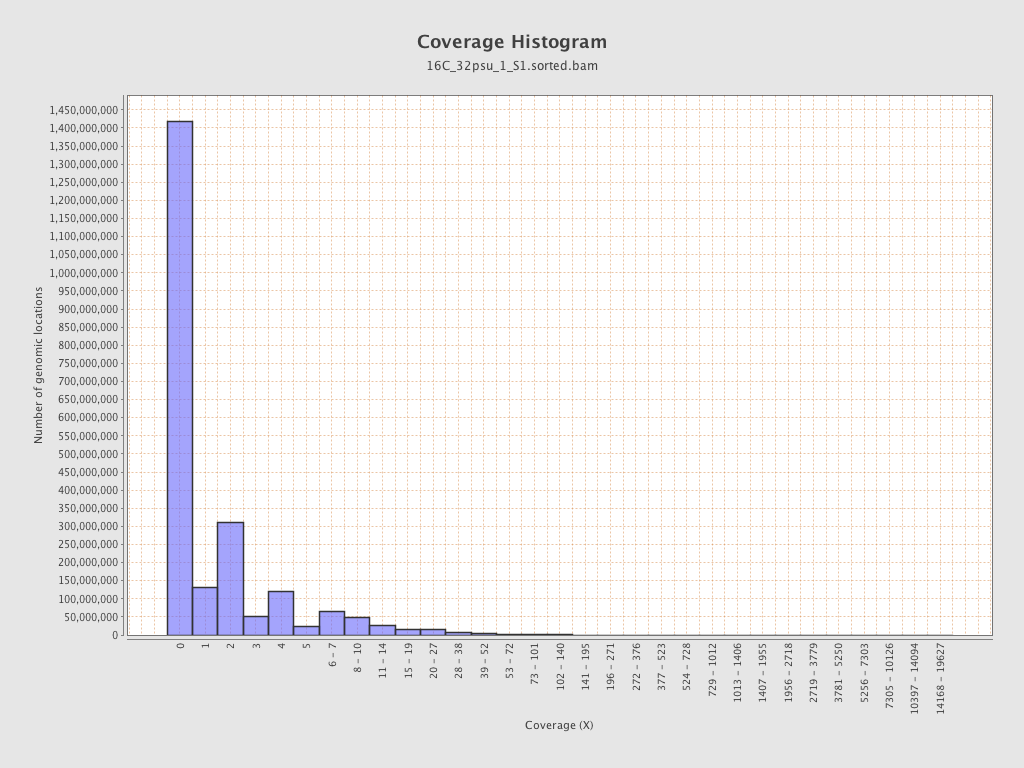
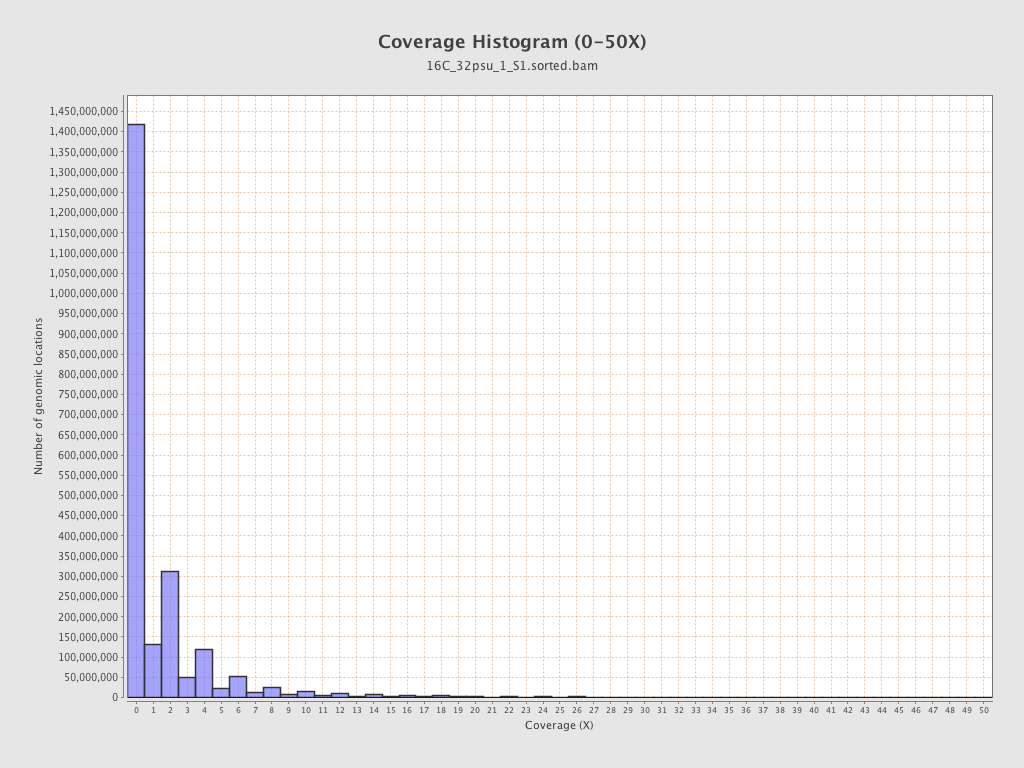
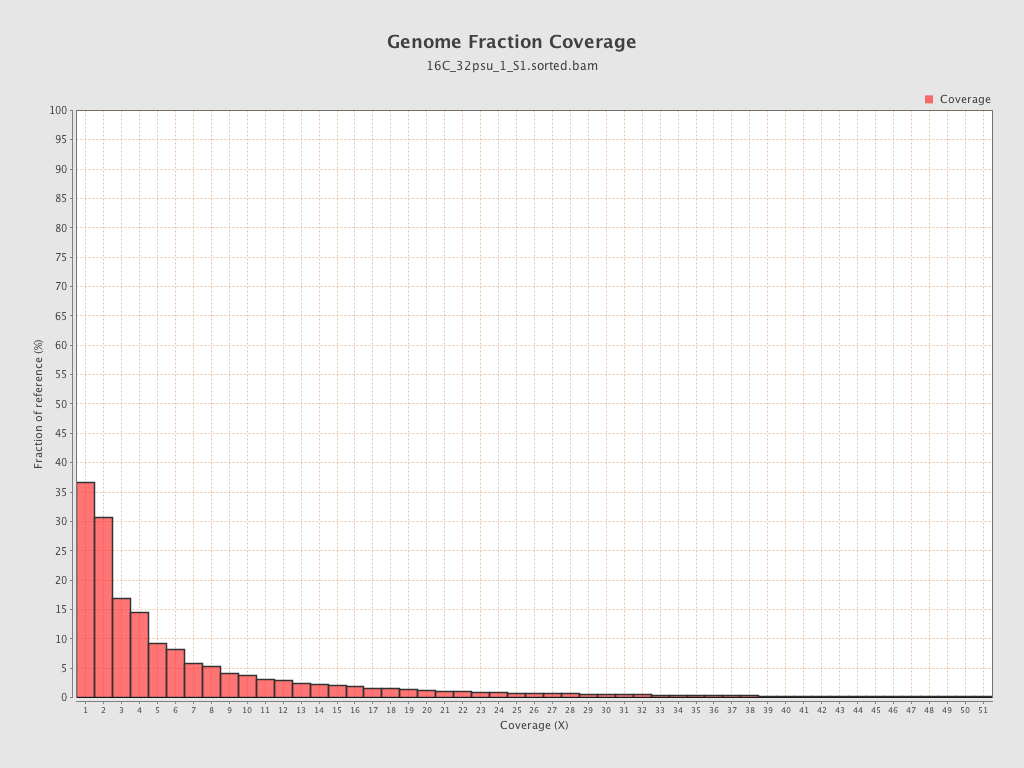
Coverage
| Mean | 1.7658 |
| Standard Deviation | 12.6447 |
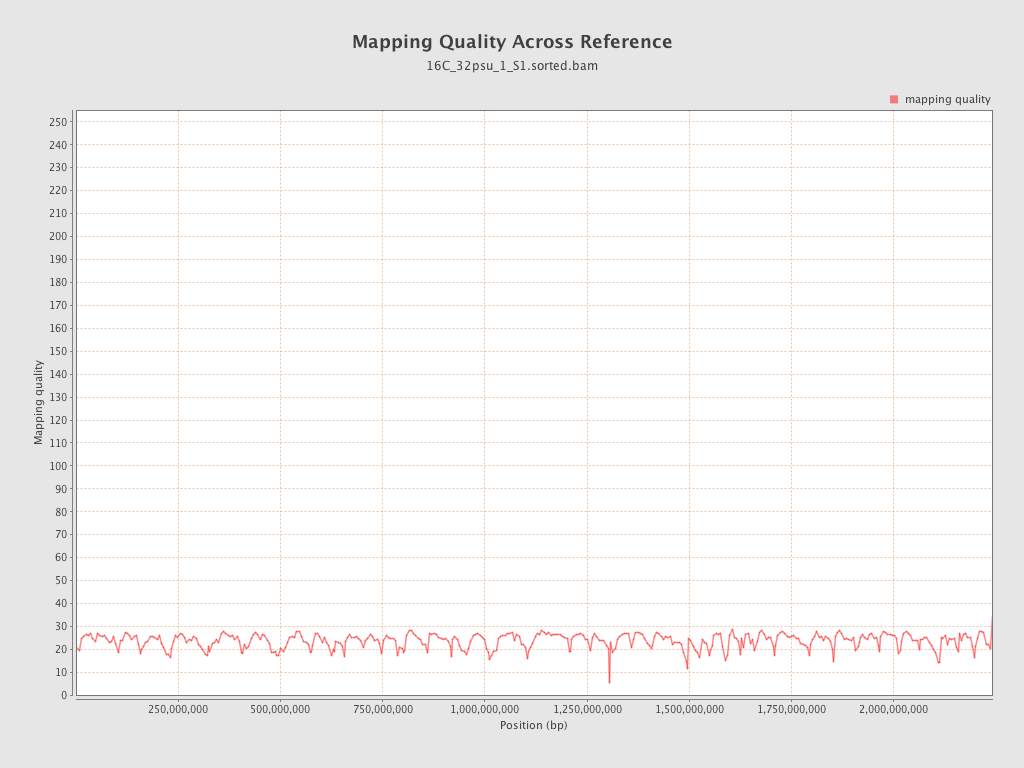
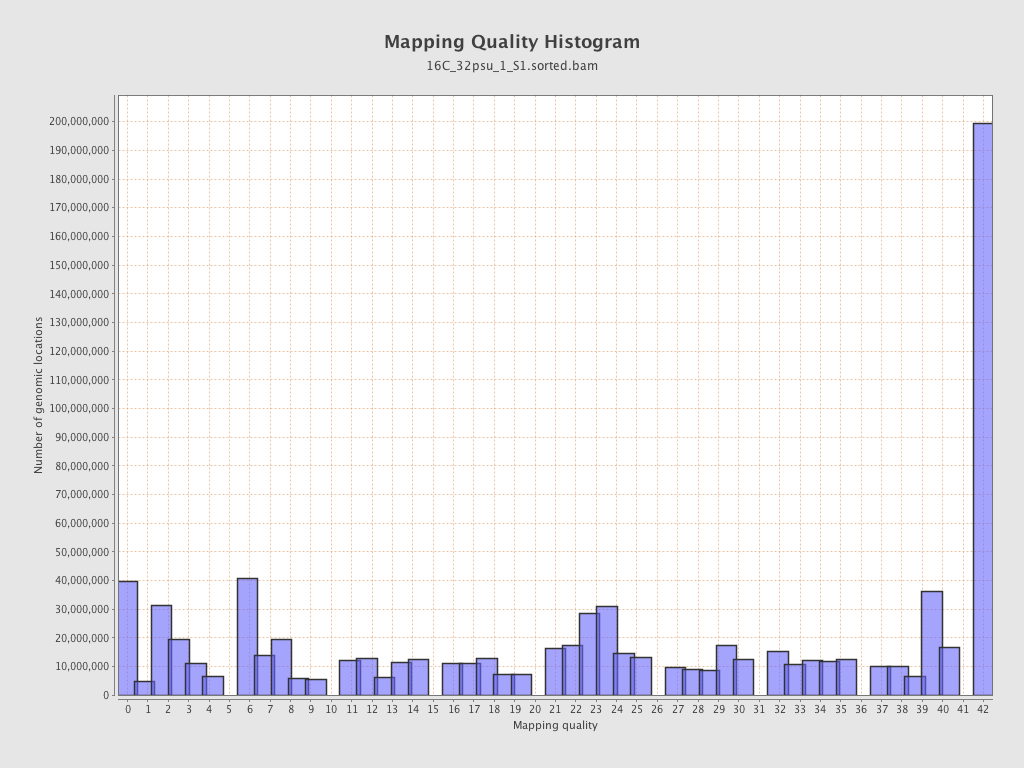
Mapping Quality
| Mean Mapping Quality | 23.55 |
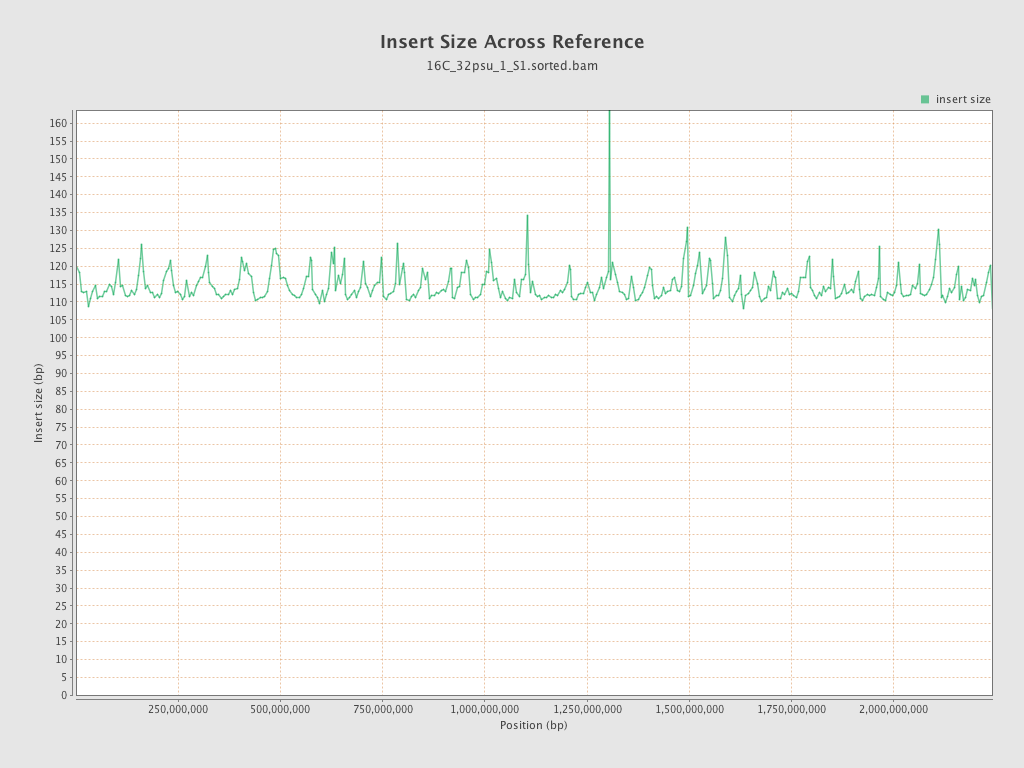
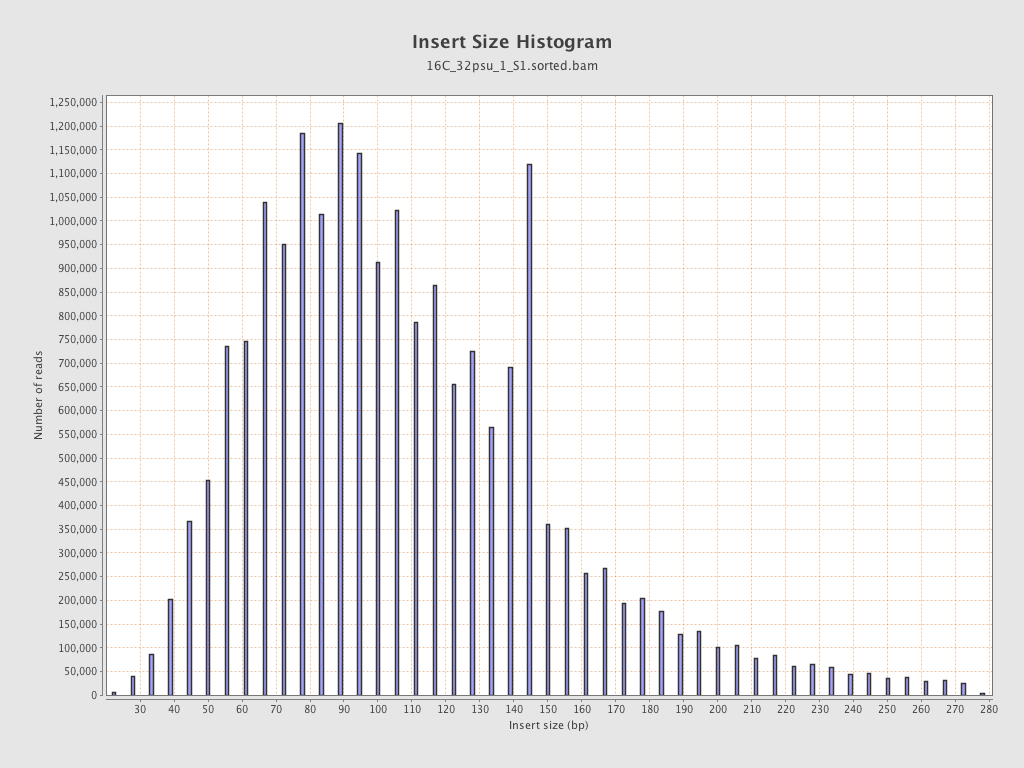
Insert size
| Mean | 114.3 |
| Standard Deviation | 52.51 |
| P25/Median/P75 | 79 / 104 / 139 |
Mismatches and indels
| General error rate | 21.24% |
| Mismatches | 810,860,900 |
| Insertions | 9,445,805 |
| Mapped reads with at least one insertion | 17.33% |
| Deletions | 6,523,318 |
| Mapped reads with at least one deletion | 14.27% |
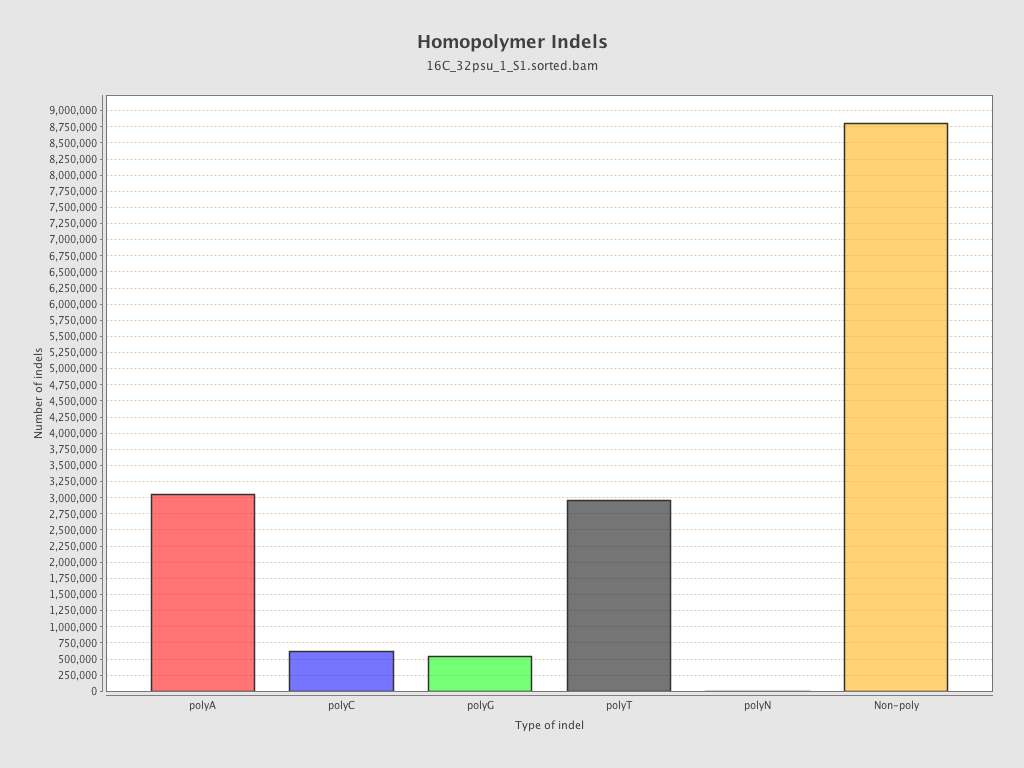
| Homopolymer indels | 44.91% |
Chromosome stats
| Name | Length | Mapped bases | Mean coverage | Standard deviation |
| NC_027300.1 | 159038749 | 292129121 | 1.8368 | 10.243 |
| NC_027301.1 | 72943711 | 120848732 | 1.6567 | 6.3189 |
| NC_027302.1 | 92503428 | 156260445 | 1.6892 | 5.5213 |
| NC_027303.1 | 82398023 | 148823154 | 1.8061 | 9.4989 |
| NC_027304.1 | 80503876 | 136262231 | 1.6926 | 8.5255 |
| NC_027305.1 | 87043187 | 144898593 | 1.6647 | 13.2493 |
| NC_027306.1 | 58785265 | 102545331 | 1.7444 | 12.0587 |
| NC_027307.1 | 26434011 | 46872759 | 1.7732 | 17.6759 |
| NC_027308.1 | 141712163 | 247543278 | 1.7468 | 9.3396 |
| NC_027309.1 | 116138521 | 215585168 | 1.8563 | 20.0022 |
| NC_027310.1 | 93888508 | 160786573 | 1.7125 | 8.4947 |
| NC_027311.1 | 91880962 | 165966402 | 1.8063 | 13.9231 |
| NC_027312.1 | 107758822 | 197160546 | 1.8296 | 8.1422 |
| NC_027313.1 | 93901823 | 165740920 | 1.765 | 10.0303 |
| NC_027314.1 | 103963436 | 179108555 | 1.7228 | 5.837 |
| NC_027315.1 | 87796322 | 158221745 | 1.8021 | 8.8871 |
| NC_027316.1 | 57682537 | 95529800 | 1.6561 | 8.1938 |
| NC_027317.1 | 70695409 | 127631743 | 1.8054 | 10.7999 |
| NC_027318.1 | 82978132 | 148308273 | 1.7873 | 26.3235 |
| NC_027319.1 | 86795997 | 154942186 | 1.7851 | 9.5425 |
| NC_027320.1 | 58021487 | 99312774 | 1.7117 | 5.7976 |
| NC_027321.1 | 63420196 | 121904392 | 1.9222 | 25.5632 |
| NC_027322.1 | 49854004 | 90975648 | 1.8248 | 8.7326 |
| NC_027323.1 | 48650976 | 80784316 | 1.6605 | 6.3436 |
| NC_027324.1 | 51481326 | 90251939 | 1.7531 | 8.8129 |
| NC_027325.1 | 47900953 | 77337302 | 1.6145 | 6.9861 |
| NC_027326.1 | 43943985 | 77203360 | 1.7569 | 8.1257 |
| NC_027327.1 | 39600944 | 71107246 | 1.7956 | 12.2343 |
| NC_027328.1 | 42488238 | 68765866 | 1.6185 | 9.3283 |
| NC_001960.1 | 16665 | 12875273 | 772.5936 | 1,212.822 |